Asking and Answering Fundamental Questions
We explore biological phenomena at the molecular and mechanistic levels to make discoveries about life and health.
Scientists in the Department of Biological Chemistry integrate many technologies and approaches to study the molecular processes that are the foundation for life and health. Research projects can be grouped into the four broad areas of Biochemical Signaling, Macromolecular Structure & Mechanism, Protein Processing & Folding and Regulation of Gene Expression.
Close ties and cross-appointments with the Life Sciences Institute, the Michigan Neuroscience Institute, the Chemical Biology Program, the LSA departments of Biology, Biophysics, and Chemistry, as well as fellow Medical School departments, position Biological Chemistry at the center of a strong and diverse biochemical community at the University of Michigan.
Our interdisciplinary research into the molecular mechanisms of life offers a wide variety of investigative opportunities to students, fellows, and scientific collaborators.

Probe how cells respond to changes in their environment.
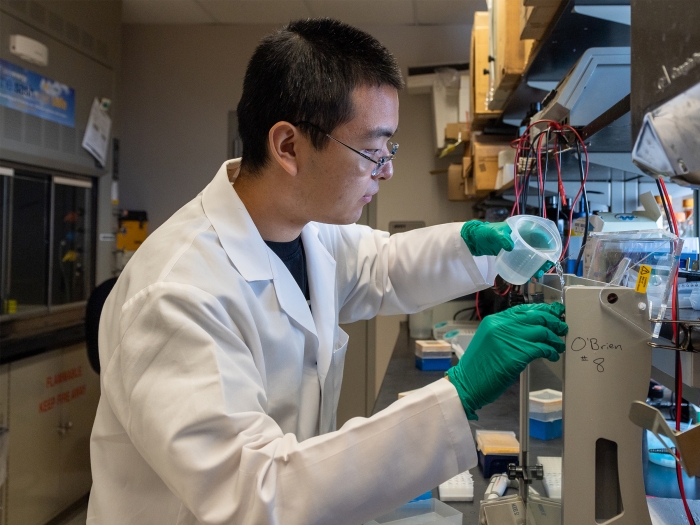

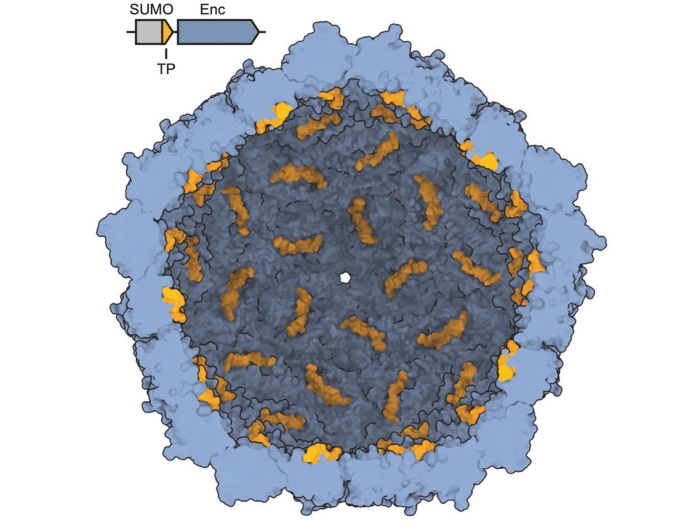
Focus in on how biological macromolecules function at the molecular and atomic level.
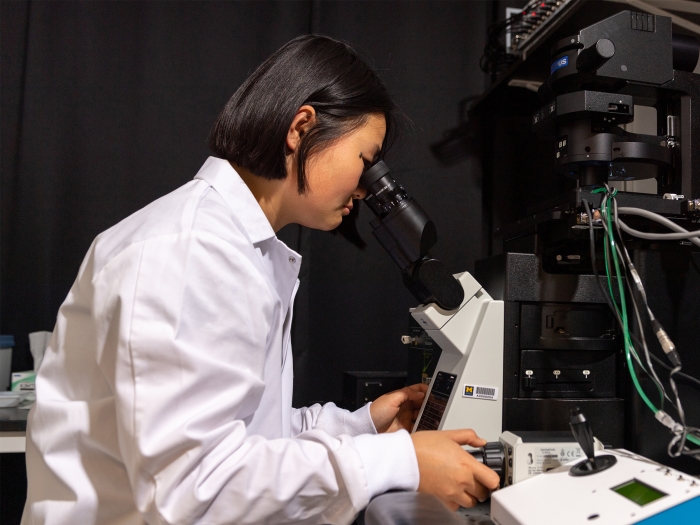

Understanding the folding, translocation, and processing of proteins is integral to modern molecular and cell biology.
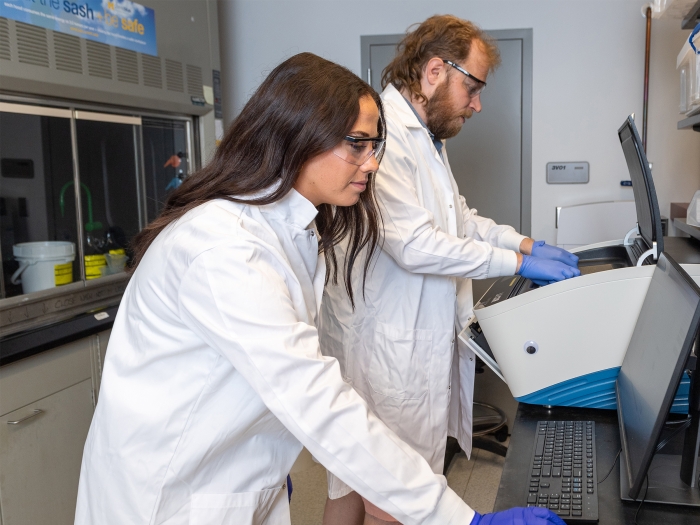

Use and develop groundbreaking research techniques to uncover the mechanisms that govern gene expression.

Patrick O'Brien Lab

Uhn-Soo Cho Lab

James Morrissey Lab

Patrick O'Brien Lab

Chase Weidmann Lab

Yan Zhang Lab

Stephen Ragsdale Lab

Raymond Trievel Lab

James Morrissey Lab

Lydia Freddolino Lab

Ruma Banerjee Lab

Lydia Freddolino Lab

Uhn-Soo Cho Lab

Rachel Niederer Lab

Ryan Baldridge Lab

Ruma Banerjee Lab

Yan Zhang Lab