A team from the University of Michigan Department of Computational Medicine and Bioinformatics (DCMB) received funding from the NIH Common Fund’s Somatic Mosaicism across Human Tissues (SMaHT) Network. The grant is for a two-part Technology and Tool Development (TTD) Project with immediate funding of nearly $800,000 to launch the first part of the project. With its second phase, the project funding will total $2.7 million. By joining this consortium, DCMB co-PIs Ryan Mills, Ph.D., and Alan Boyle, Ph.D., will collaborate with leaders in the field of cellular biology and bioinformatics, developing state-of-the-art technologies to better detect somatic mosaicism in different cell types and tissues, and across human life stages.
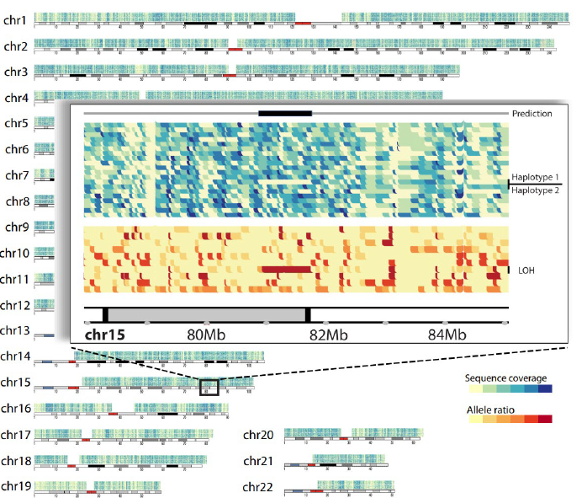
Example of a large somatic deletion identified from whole genome single cell DNA sequencing using Oxford Nanopore Technologies, as denoted by yellow (top) and red (bottom) rectangles in individual cells (rows).
Our tissues are made of a multitude of cells, called somatic cells (as opposed to reproductive cells) that replicate themselves. As they duplicate, most of our somatic cells remain genetically the same, but some of them turn out differently, creating a so-called “somatic mosaicism.” This somatic genetic variation occurs when some of our cells accumulate DNA changes over time. Somatic mosaicism can alter how cells function and may influence human development, disease, aging, and other physiological measures across the lifespan. However, to this day, deep investigations into the biology of this cellular differentiation have been challenged by the limits of both molecular assay techniques and computational methodology and capacity. To boost this area of research, NIH is launching the SMaHT Network, supported by $140 million in funding over five years. The network gathers experts across many institutions and disciplines, ranging from cellular biology to bioinformatics.
The DCMB team is well positioned to make a substantive contribution to the network.
Collaborators in the Mills and Boyle labs have extensive collective experience developing tools for identifying somatic mosaicism in the human brain, including recent surveys of single-nucleotide variants (SNV) prevalence from whole genome and exome sequencing, copy number variations (CNV) from single cell short-read and nanopore genome sequencing, and retrotransposons through targeted capture. With their SMaHT Network contribution, the team will seek to improve, optimize, and extend its approaches to other human tissues.
“The SMaHT Network will provide an excellent platform for systematically identifying and cataloging natural human somatic mosaicism across tissues and will provide a resource for future studies to explore their associations with aging and age-related diseases,” said Mills.
Working in both molecular and computational labs
With this NIH support and the SMaHT Network, Mills and Boyle will improve molecular assays for targeted bulk capture and single cell nanopore sequencing. They will demonstrate that these techniques, previously developed for brain tissue studies, are scalable, both through multiplexed barcoding and adaptive targeted sequencing. They will then expand their work to apply these techniques across different tissues, according to the SMaHT Network’s priorities.
"Our recent methodological advancements have put us in a unique position to contribute to the SMaHT consortium, paving the way for research breakthroughs in the understanding of somatic mosaicism and unlocking new avenues for scientific exploration", said Boyle.
In the computational lab, they will develop algorithms for detecting somatic mosaicism from single cell and bulk tissue data by adapting and improving their existing methods for detecting SNVs, CNVs, MEIs, and STRs from single cell and targeted capture nanopore data. They will demonstrate that these tailored approaches outperform typical germline or clonal based approaches and are applicable across the variant allele frequency spectrum. Once they have determined the best practices for somatic mutation detection, they will apply these to study SMaHT prioritized tissues.
Overall, the DCMB team will provide efficient, scalable, and documented software packages for their computational approaches, with extensions for cloud computing as necessary. The team plans to deploy its software to open-source software repositories for ease of installation and application. They will work with consortium efforts to integrate their approaches into standardized analysis pipelines for large-scale application across project data, including incorporating project-defined output formats into their software.
The DCMB team will particularly collaborate with Dr. Michael McConnell at the Lieber Institute for Brain Development, who is also a principal investigator on the award.
This research is supported by the NIH Common Fund grant 1 UG3 NS132084-01.
Congratulations to Drs. Mills, Boyle, and McConnell, and their teams!

Alan Boyle, Ph.D.

