Our genome is intricately structured at multiple levels in the cell. However, how these structural aspects affect cellular function and cause disease is largely unknown. HG faculty members are uncovering these mysteries of our genome by employing cutting edge experimental and computational approaches. Our work encompasses the realms of basic, applied and clinical research.
Variation in our DNA essentially makes us different from one another. A major focus of our research is to identify and map these sequence variations in humans and other species, and explore its impact on diversity and evolution (Kidd, Kitzman, Mills). We resolve different types of genomic variation and integrate this information with medical data to better understand human health and disease (Mills). We also investigate molecular mechanisms driving de novo variants in germ cells (Lukaszewicz). We aim to understand how ‘jumping genes’ or retrotransposons affect the structure our genome function (Moran).
Our DNA is susceptible to damage which when not properly contained by the cell’s inherent repair mechanisms can lead to cancers and developmental diseases. We are closely investigating these repair mechanisms, which could then pave the way towards designing efficient disease therapies (Cheung, Glover, Sekiguchi, Wilson).
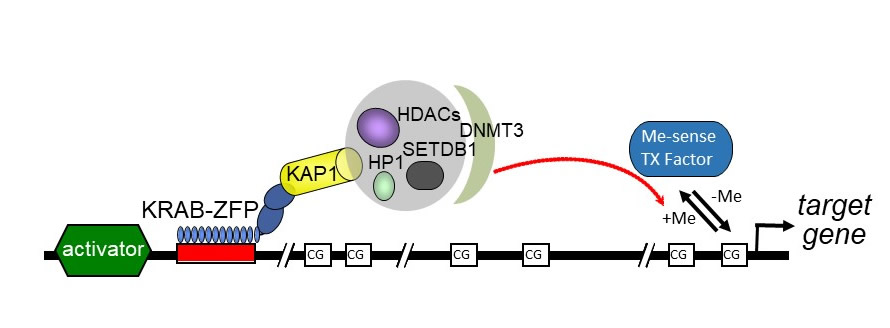
Another aspect to our research is to identify genes involved in specific cellular processes and characterize what factors and mechanisms regulate their expression (Antonellis, Cheung, Robins). We are testing hypotheses to determine the mechanisms of post-transcriptional gene regulation in stress, aging, and neurodegeneration using molecular genetics and microscopy approaches (Moon). We are studying the process of spermatogenesis in the context of male infertility, and our work is unraveling the complex genomic architectures of the sex chromosomes (Hammoud, Lukaszewicz, Muller).
Our faculty members are at the forefront of technological advancements and breakthroughs in research. We employ novel single cell genomics and super resolution microscopy methods (Camper, Hammoud). We are tackling research problems in unprecedented throughput by developing techniques such as nascent RNA sequencing (Wilson), massively parallel mutagenesis and transcriptional assays along with big data analysis tools (Boyle, Kitzman, Parker).