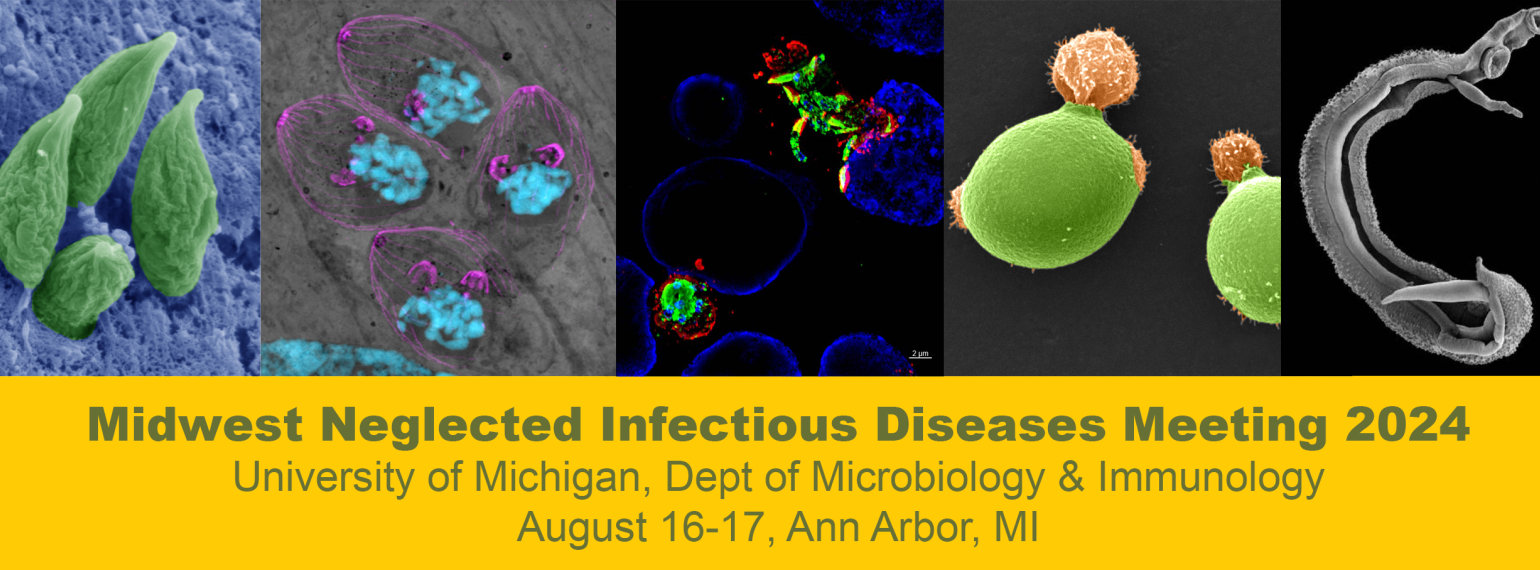
All the wonderful things
that happened in M&I this year!
 Welcome to the Department of Microbiology and Immunology at the University of Michigan
Welcome to the Department of Microbiology and Immunology at the University of Michigan
We are an inclusive community whose passion for discovery, service, leadership and sharing of knowledge addresses global microbiology and immunology challenges
Follow us on Twitter @UMMicroImmuno!
Please click on the navigation links on the left hand side of the page to open several more menu items to help acquaint you with our department, our university and our town!
We are training the next generation of scientists.
In Microbiology & Immunology, graduate students and postdoctoral fellows work side by side with principal investigators to test hypotheses of medical importance. Laboratory training is supplemented with rigorous coursework, seminar series, journal clubs, and presentation of research findings at national meetings. Learn more about our excellent education programs.
Helping unlock the mysteries of the microbial world and the immune system.
Our faculty study many aspects of microbial pathogenesis, microbial communities and the interactions with and functions of the immune system. Learn more about our areas of research.